SAN DIEGO—During the past several decades, an understanding of the pathogenesis of many rheumatic diseases, such as rheumatoid arthritis (RA) and psoriatic arthritis (PsA), has been illuminated by the discovery of the key role played by certain human leukocyte antigen (HLA) alleles. What is now known, according to Robert Winchester, MD, professor of pediatrics, medicine, and pathology in the department of pathology and cell biology, division of rheumatology, at the College of Physicians and Surgeons of Columbia University in New York, is that both RA and PsA are autoimmune in pathogenesis, and that the most important determinants of disease susceptibility are certain alleles encoding the particular major histocompatibility complex, HLA. “While the roles and significance of the HLA genes differ in these two diseases, the HLA molecules encoded by these genes likely contribute importantly to specifying the type of adaptive immune response that underlies each disease,” he said.
What are the implications of this knowledge on understanding the pathogenesis of RA and PsA? Dr. Winchester walked participants through an answer to this question via an elegant lecture on the meaning and significance of HLA associations in rheumatologic diseases during the 2013 Paul Klemperer, MD, Memorial Lecture, “Rheumatoid Arthritis, Psoriatic Arthritis and Autoimmunity: Good Genes, Elegant Mechanisms, Bad Results,” here at the 2013 ACR/ARHP Annual Meeting, held October 26–30. [Editor’s Note: This session was recorded and is available via ACR SessionSelect at www.rheumatology.org/sessionselect.]
Adaptive Immune Response
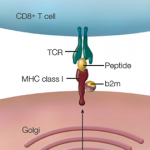
The heart of the adaptive immune response is the T-cell receptor–peptide–HLA molecule interaction, he said, which is also the cornerstone on which the development of autoimmunity rests. Autoimmunity develops through two major consequences that occur when the T cells of the adaptive immune system are challenged to recognize many different pathogen peptides. The first consequence, he said, is somatic generation and thymic selection of T-cell repertoire on self-peptides bound to self-HLA. “Because the entire T-cell portion of the adaptive immune system is built on physiologic self-recognition, this leads to the risk that certain individuals will develop autoimmunity,” he said.
The second consequence is diversification of the HLA system to develop new alleles that better bind these various peptides because of the need of HLA molecules to bind and anchor each of the different pathogen peptides with high affinity, he said. “Since certain HLA molecules bind particular self-peptides that are expressed on cells and tissues comprising the joint, these self-peptides become targets of the autoimmune adaptive immune response,” he said, adding that, “these, in turn, drive the inflammatory response once they are triggered by still-unknown processes.”
Disease Susceptibility: Rheumatoid Arthritis
Citing work done in concert with Peter Gregersen, MD, a geneticist who heads the Robert S. Boas Center for Genomics and Human Genetics at the Feinstein Institute for Medical Research in Manhasset, N.Y., he described what is known about disease susceptibility in RA. Their research shows that RA disease susceptibility is associated with at least eight different alleles of HLA-DR locus, and that each allele encodes a similar sequence (i.e., shared epitope). According to Dr. Winchester, the alleles in patients with RA that are responsible for RA susceptibility are identical in sequence to the alleles in healthy persons (i.e., people with good genes) and are not rare abnormal genes. He said that studies show that at the genotypic level, the risk of RA is additive. For example, the risk conferred by two shared epitope alleles is greater than that conferred by one shared epitope allele. Similarly, the risk conferred by one shared epitope allele is greater than that conferred by alleles lacking the shared epitope structure, although dominant protection is not conferred by any shared epitope negative alleles. “Importantly,” he said, “the clinical phenotype of rheumatoid arthritis is essentially the same regardless of which among the eight alleles a person inherits.”
What remains unknown and critical to understanding the development of RA is the event associated with the binding of peptides influenced by the P4 pocket, he said, pointing out the limitations and problems with the hypothesis put forth by Hill and colleagues in 2003 on the conversion of arginine to citrulline that permits binding of peptides.1
Disease Susceptibility in PsA
Unlike RA, the genotype of PsA is heterogeneous, said Dr. Winchester, citing data that show that the HLA associations predisposing a person to PsA are largely different than those predisposing a person to cutaneous psoriasis. Data show that 87% of people with PsA have one or more of three susceptibility alleles (HLA-B*08, HLA-B*27, and HLA-C*06:02), none of which have common structure features, whereas the genotype associated with cutaneous psoriasis is primarily associated with HLA-C*06:02. He also said data show that phenotypes found in people with PsA differ with each of the three different genotypes, citing data that show, for example, that highly penetrant early-onset psoriasis and late-onset low-penetrant arthritis is characterized by HLA-C*06:02 and that early-onset arthritis that develops more synchronously with the onset of skin disease is more characteristic of HLA-B*27.
Report Card: Where Do We Stand
Dr. Winchester concluded by talking about the status of the progress on what is known about the HLA associations in rheumatic diseases. He emphasized that, because of the differences in the way the HLA molecules confer susceptibility to RA and PsA, progress in understanding these mechanisms is at the heart of the challenge of thwarting the processes involving interactions between HLA molecules, self-peptides, and T-cell receptors that result in disease. Saying that the general details of the HLA component are beginning to be identified, he emphasized that what remains unknown are identification of the peptides that drive the autoimmune process as well as identification and recognition of the properties of the T-cell clonotypes that drive the autoimmune processes.
“Powerful new emerging technologies give the promise that, over the next decade, understanding will be greatly advanced and likely will lead to rational therapies directed at the fundamental events driving these diseases,” he said.
Mary Beth Nierengarten is a freelance medical journalist based in St. Paul, Minn.