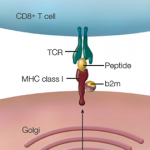
The National Academy of Sciences has convened a panel of experts to assess current regulations and recommend what additional oversight will be needed with regard to gene manipulation.
Image Credit: Sebastian Kaulitzki/shutterstock.com
Every so often, a major scientific breakthrough profoundly alters the trajectory of scientific research. In the 1960s, microbiologists sparked the recombinant-DNA revolution with the discovery that bacteria have innate immune systems based on restriction enzymes. These enzymes bind and cut invading viral genomes at specific short sequences, and scientists rapidly repurposed them to cut and paste DNA in vitro, transforming biologic science and giving rise to the biotechnology industry.1
In the 1970s, the seminal efforts of the late Nobel laureates, César Milstein, PhD, and Georges Köhler, PhD, in the Laboratory of Molecular Biology in Cambridge, UK, demonstrated the efficacy of monoclonal antibodies as probes to investigate the pathological pathway of neurological conditions. Although limited in scope, this initial application paved the way for the adoption of monoclonal antibodies as diagnostic tools. Perhaps more importantly, other investigators recognized their potential in the therapeutic sphere. Within years of their discovery, monoclonal antibodies served as an invaluable platform in the development of biologic therapies for rheumatic diseases.2 (See Rheuminations; January 2012, “A Short History of RA Therapeutics.”)
In the 1980s, using the polymerase chain reaction (PCR), we learned how to speedily amplify specific DNA sequences, creating a way to generate large amounts of genetic material from a single cell, a previously laborious technique.
More recently, a fascinating discovery that allows for the easy manipulation of an organism’s genome has rocked the world of biology. Unlike some of the other scientific breakthroughs described earlier, this one has the potential to alter our genomes, and so the use or misuse of this technology will have profound implications for society.
CRISPR: From the Yogurt Cup to the Human Genome
More than 30 years ago, scientists observed a strange pattern in some bacterial genomes: One DNA sequence would be repeated over and over again, while unique sequences would be found in between the repeats. This odd configuration, coined clustered regularly interspaced short palindromic repeats (CRISPR), was subsequently found to serve as a naturally occurring, ancient defense mechanism in a wide range of bacteria. How does it work?3
CRISPR, in combination with a set of enzymes called Cas9 (CRISPR-associated proteins), uses a guide RNA molecule to home in on its target DNA, then edits it by inserting desired sequences or disrupts it altogether. To use an analogy, these lab-made CRISPR systems are the cellular version of a word processor’s find-and-replace function.