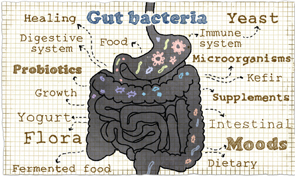
Recent data suggest that our commensal microbiota are not passive, but reflect and even induce systemic disease.
T. L. Furrer/shutterstock.com
The microbiome comprises diverse microbial flora, including bacteria, viruses and fungi, that live on mucosal surfaces, predominantly the skin and digestive tract. Microbes evolved billions of years prior to the development of modern Homo sapiens 200,000 years ago; we have always existed with their ubiquitous presence. Despite this, the first microbe was not visualized until 1665 when Robert Hooke and Antoni van Leeuwenhoek published a drawing of Mucor.1 Alexander Fleming’s discovery of penicillin in 1928 not only revolutionized modern medicine, but also established the understanding of bacteria as single agents primarily antagonistic to human health. Microbial research focused on attacking and eliminating bacteria and viruses, with little regard for the commensal flora that live inside of us.
These residents were thought to remain relatively static, existing independently of host health or disease. Recent data suggest that our commensal microbiota are not passive, but reflect and even induce systemic disease. Advances in technology and bioinformatics over the past decade have stimulated this expanding field, allowing for more comprehensive analysis of this complex ecosystem, and now not only encompass the microbiome, but also the proteome and metabolome, the proteins and metabolites these flora produce. Most research has focused on bacterial flora, but there are increasing studies investigating the virome and fungome. Growing data suggest that alterations in this flora, termed dysbiosis, play a large role in the development of rheumatic diseases.
Rheumatoid Arthritis (RA)
RA has a concordance of 15.4% in monozygotic twins, strongly suggesting a significant environmental role in disease pathogenesis.2 There has been long-standing suspicion that RA is secondary to infection. A review article in 1931 noted that the popular term for RA at the time was chronic infectious arthritis and the underlying etiology of an infection was the most accepted hypothesis for disease pathogenesis.3 For more than 65 years, treatment of RA has utilized antimicrobials; sulfasalazine contains sulfapyridine, an antibiotic, and randomized trials have shown minocycline and macrolides are effective at decreasing disease activity.4
However, data demonstrating a microbial role in RA pathogenesis remain indirect. Examination of fecal, salivary and dental microbiomes of patients with established RA demonstrates that dysbiosis in the oral and gastrointestinal tract correlates with reduced disease activity and resolves with treatment.5 The association between periodontal disease caused by Porphyromonas gingivalis and patients with RA who have antibodies to citrullinated proteins (ACPA) proposes that mucosal inflammation caused by P. gingivalis can lead to the generation of ACPA and subsequently RA.6-8