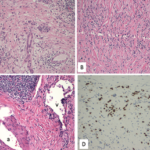
IgG4-related disease (IgG4-RD) often presents with elevated levels of serum IgG4, as well as diffuse swelling, mass formation and/or fibrosis of affected organs. Patients with IgG4 have genetic risk factors and an innate immune response to pathogens that mimic self-antigens. These patients are also noteworthy because their pathophysiology includes active involvement of Th2 and T follicular helper (Tfh) cells.
IgG4-related disease (IgG4-RD) often presents with elevated levels of serum IgG4, as well as diffuse swelling, mass formation and/or fibrosis of affected organs. Patients with IgG4 have genetic risk factors and an innate immune response to pathogens that mimic self-antigens. These patients are also noteworthy because their pathophysiology includes active involvement of Th2 and T follicular helper (Tfh) cells.
New research suggests that prednisone treatment can selectively modulate the signatures of regulatory T (Treg) cells, eosinophils and neutrophils in these patients. Brandon W. Higgs, PhD, lead bioinformatics scientist at MedImmune in Gaithersburg, Md., and colleagues published their molecular profile data online Dec. 14 in Scientific Reports.1 The investigators used whole transcriptomic sequencing to identify and distinguish both cell- and pathway-associated activation in patients with IgG4-RD. They also characterized the molecular differences and effects from prednisone treatment. Their study included patients with salivary gland lesions (RD-SG), patients without salivary gland lesions (RD-nonSG) and patients with IgG4-related retroperitoneal fibrosis (RF).
The investigators used RNA sequencing to examine a cohort of RD-SG (n=25), RD-nonSG (n=11) and RF (n=3) patients and compared them with controls (n=10). Specifically, they used principal components analysis (PCA) to characterize the transcriptome profile of the three diseases relative to healthy controls and found patients with RD—both SG and nonSG—overexpressed IgG4 and IgE. The team then constructed a linear model that allowed them to identify the genes across the transcriptome that most correlated with levels of expression of IgG4 mRNA. They found Treg, Th2, eosinophil and neutrophil gene signatures were all overexpressed in patients with RD-SG and RD-non-SG. In contrast, the B cell signature was suppressed in these patients. When the researchers looked more closely, they found genes associated with mitosis, cell cycle and replication were the ones most likely to be correlated with IgG4 expression. Specifically, IgG4, IgE and cell-specific signatures were all regulated in the patients, which suggested to the authors that IgG4-related disease may be the result of an imbalance of immune and inflammatory cells.
The researchers then looked more closely at the effects of prednisone on IgG4-RD. Within their group of patients, eight RD-nonSG and 12 RD-SG patients received prednisone treatment and/or glucocorticoid-sparing agents. Additionally, six patients with RD had a longitudinal timepoint. A heatmap of the molecular gene signatures revealed that prednisone treatment affected all T cell sub-populations. Specifically, prednisone treatment was associated with increased neutrophil, but decreased Treg signatures. When the investigators compared the patients at baseline with their status during a flare, they found no association with induction of Th1, Th2, Treg and eosinophil gene signatures. However, they did find an inverse relationship between the flare and both the B cell signature and Tfh genes.
In their discussion the authors write, “We show the importance of the T and B cell axis with molecular profiling across RD-SG, RD-nonSG and RF, as well as features that distinguish these three diseases. Future work seeks to better understand the molecular mechanisms at relapse or recurrence following steroid reduction in these patients.”
Lara C. Pullen, PhD, is a medical writer based in the Chicago area.
Reference
- Higgs BW, Liu Y, Guo J, et al. High-throughput RNA sequencing reveals distinct gene signatures in active IgG4-related disease. Sci Rep. 2017 Dec 14;7(1):17567. doi: 10.1038/s41598-017-17602-9.